Metagear: an R tool to systematic reviews and meta-analyses

Systematic reviews are a type of review that uses repeatable analytical methods to collect and analyse secondary data. When done properly, systematic reviews are an excellent way to synthesise the results from many studies. Systematic reviews are often paired with meta-analyses: statistical analyses that combine the results of multiple scientific studies that address the same (or similar) questions. For example, we might want to determine whether transcranial direct current stimulation improves cognitive ability in people who have suffered a stroke (1). If this was the case, we would first conduct a systematic review to identify published and unpublished scientific papers and other documents such as theses and reports, often referred to as grey literature, that have used transcranial direct current stimulation as an intervention to improve cognitive abilities in stroke survivors. We would then use meta-analytical approaches to combine the results from these studies to obtain a more precise estimate of the true effect of transcranial direct stimulation.
Covid-19 has presented many challenges for researchers. For some, the inability to conduct research in the clinical or laboratory setting has led them to search for other research options, including systematic reviews and meta-analyses. While this seems sensible, systematic reviews and meta-analyses have a long, rich history. They are invaluable when done properly, but unhelpful when done improperly. While there are several good books on the topic, it can be difficult for newcomers to conduct high-quality systematic reviews. My personal advice is to find experts who have previously published systematic reviews and meta-analyses to collaborate with, that is what I did (thank you Jo and Georgia!). 🙂
Tools for systematic reviews and meta-analysis
An important decision we have to make when we start our systematic review relates to our choice of tools. A systematic review has several steps, some of which can be automated, some of which can be semi-automated, and some of which must be done manually. Importantly, we must keep track of our entire work flow and the various decisions we have made along the way. One option is to use commercial software. This is a good option is your institute has a license; if not, you will be paying USD240 per year for one systematic review (a great motivation to get your review done in one year!). However, such tools often involve a web-based work flow that requires an internet connection with lots of pointing and clicking. Given that systematic reviews are often completed by teams of researchers and students, the ability to have multiple users work simultaneously and remotely is one of the strengths of such systems.
However, being the person I am, when I got involved in a systematic review I immediately started to look for open-source alternatives. An initial search did not uncover any viable options, thus our team started to code our own tools and work flow in Python (more on this in a later post, possibly).
METAGEAR
However, in the intervening period, I came across a great little project by Marc Lajeunesse called METAGEAR.
METAGEAR is an package for the R statistical programming language.
Dr. Lajeunesse wrote a paper on the package in 2016,
and has since started a great Youtube channel with loads of courses and tutorials dedicated to teaching various aspects of systematic reviews.
As stated in the paper:
"METAGEAR is a comprehensive, multifunctional toolbox with capabilities aimed to cover much of the research synthesis taxonomy: from applying a systematic review approach to objectively assemble and screen the literature, to extracting data from studies, and to finally summarize and analyse these data with the statistics of meta-analysis.
Current functionalities of metagear include the following: an abstract screener GUI to efficiently sieve bibliographic information from large numbers of candidate studies; tools to assign screening effort across multiple collaborators/reviewers and to assess inter-reviewer reliability using kappa statistics; PDF downloader to automate the retrieval of journal articles from online data bases; automated data extractions from scatter-plots, box-plots and bar-plots; PRISMA flow diagrams; simple imputation tools to fill gaps in incomplete or missing study parameters; generation of random-effects sizes for Hedges' d, log response ratio, odds ratio and correlation coefficients for Monte Carlo experiments; covariance equations for modelling dependencies among multiple effect sizes (e.g. with a common control, phylogenetic correlations); and finally, summaries that replicate analyses and outputs from widely used but no longer updated meta-analysis software."
As stated above, METAGEAR has many useful tools to help researchers as they traverse the various steps of a systematic review and meta-analysis. These include a simple user-interface to screen abstracts and titles:

tools to generate a systematic review flow chart:

and tools for data extraction and analysis:

If you know the basics of R and you are embarking on your first systematic review (or would like to move away from web-based, proprietary software,
you may want to consider METAGEAR.
OpenMEE
Dr. Lajeunesse and other collaborators have also contributed OpenMEE, an intuitive, open-source software for meta-analysis in ecology and evolutionary biology.
It has a Python front-end graphical user interface that runs R in the background.

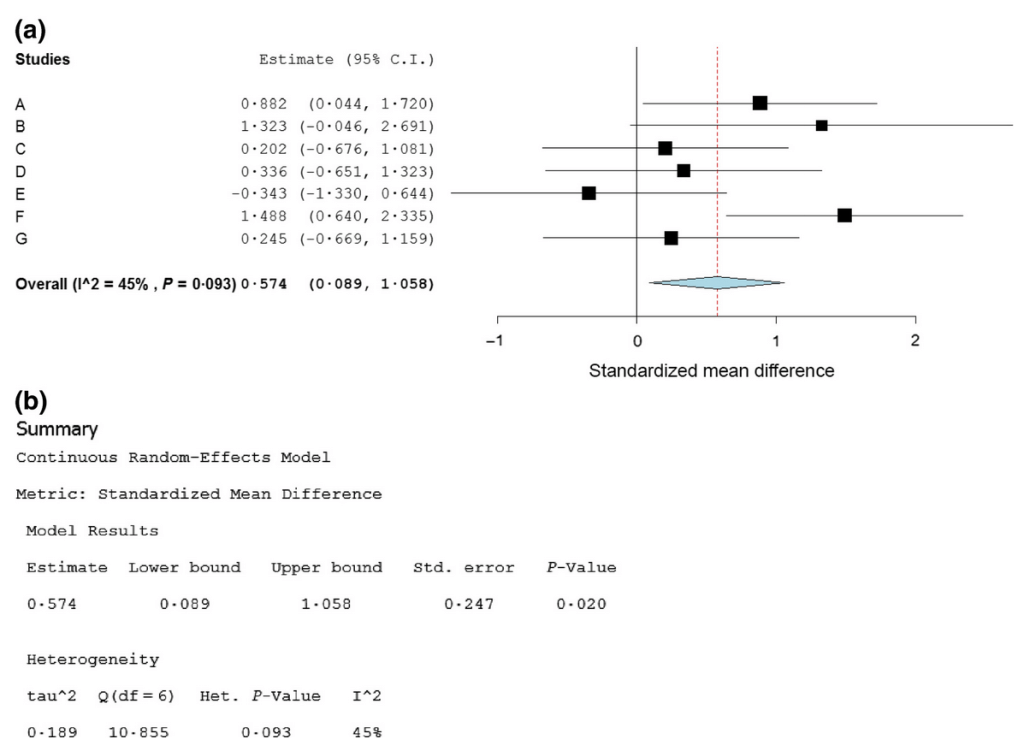
OpenMEE is another great example of great open-source software.
Conclusion
Systematic reviews and meta-analyses are incredibly useful. Given the limitations imposed on researchers by Covid-19 restrictions and the never ending pressure by Institutions for their researchers to publish more and more, many have turned to systematic reviews and meta-analyses to fill the gap. Part of delving into a new field or adopting a new approach is finding the right tools. While there are proprietary, paid options available, it is definitely worthwhile to have a look at what is available on the open-source side of things. Who knows, my team might even make our Python implementation public once we have worked out the kinks.
