R: Analysing small datasets – Part 3

In the previous post, when repeated measures data from 10 subjects in 2 conditions were compared, it seemed that subjects who took drug 1 slept fewer hours compared to when they took drug 2.
How might we test whether the median number of hours of sleep after drug 1 is less than after drug 2? We can calculate the difference between the medians for this sample. We would also like a measure of precision of the difference of the medians (95% CI) to know how much it might change if subjects in the sample change. The problem is that the median is a non-parametric statistic based on the rank of observed data points (rather than numeric values) and it does not rely on data belonging to any particular probability distribution. So we can’t use parameters or characteristics of probability distributions to calculate 95% CI about the difference of the medians.
What we could do is generate another sample and calculate a difference of the medians for that sample. If we can generate many samples with many differences of medians, it would be possible to see how the difference of the medians “jumps around” with repeated sampling, and calculate the 95% CI. The technical name for how this is done is a statistical procedure called bootstrapping.
The first time I heard about bootstrapping from my PhD supervisor, I had immediate visions of Pirates of the Caribbean (maybe to do with Bootstrap Bill Turner), and Baron Munchausen who was widely (though mistakenly) reported to have pulled himself out of a swamp by his bootstraps. Actually bootstrapping isn’t quite that. The term has to do with an automated process that can proceed without external input, like when you “boot up” your computer, the operating system then loads software as needed.
In statistical bootstrapping, we generate a new sample by sampling with replacement from the original sample and calculate an estimate of the effect for the new sample (here, we calculate the difference of the medians between drug 1 and 2). Sampling with replacement does mean that some subject values may be included multiple times in some bootstrapped samples or not at all in others. But, by generating lots and lots of samples, each with a between-condition difference of the medians, we can observe how the difference of the medians is distributed over many samples and calculate a 95% CI.
This is one way of doing it in R, as outlined here:
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 |
# install packages: comment out line below if packages are installed
install.packages('boot')
library(boot)
# create bootstrapping function using data in long format
n <- length(df_wide$extra.1)
m <- length(df_wide$extra.2)
dif.median <- function(data, i) {
d <- data[i,]
n1 <- n+1
m1 <- n+m
median(d$extra[1:n]) - median(d$extra[n1:m1]) # indexing of data in long format
}
# bootstrapping with 1000 replications
set.seed(10)
dif.boot <- boot(df, dif.median, R=1000, strata=df$group)
# view results
print(dif.boot)
plot(dif.boot)
# get 95% CI
boot.ci(dif.boot, conf=0.95, type='bca')
|
Line 1-3. Install the boot package if required and load the library.
Line 6-7. Using the data in wide format, calculate the number of observations for each of the conditions (this means we could use this procedure on observations with different lengths, in other instances).
Line 8-13. Create a function called dif.median that produces the statistic to be bootstrapped (ie. the difference of the medians), and include an indices parameter i as a function argument that the boot() function can use later to generate samples with replacement.
Line 15. Set a seed value so bootstrapped results can be consistently reproduced.
Line 16. This is where the action happens: call the boot() function, pass it the data df which is in long format, the dif.median function, and generate 1000 samples with replacement where samples are stratified by group (ie. drug). Assign the bootstrapped results to the object dif.boot, then print and plot the results and calculate the bootstrapped 95% CI.
The following output is produced:
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 |
STRATIFIED BOOTSTRAP
Call:
boot(data = df, statistic = dif.median, R = 1000, strata = df$group)
Bootstrap Statistics :
original bias std. error
t1* -1.4 -0.2168 1.222873
BOOTSTRAP CONFIDENCE INTERVAL CALCULATIONS
Based on 1000 bootstrap replicates
CALL :
boot.ci(boot.out = dif.boot, conf = 0.95, type = "bca")
Intervals :
Level BCa
95% (-4.29, 0.75 )
Calculations and Intervals on Original Scale
|
Plotting the bootstrapped results produces the following Figure 1:
Figure 1:
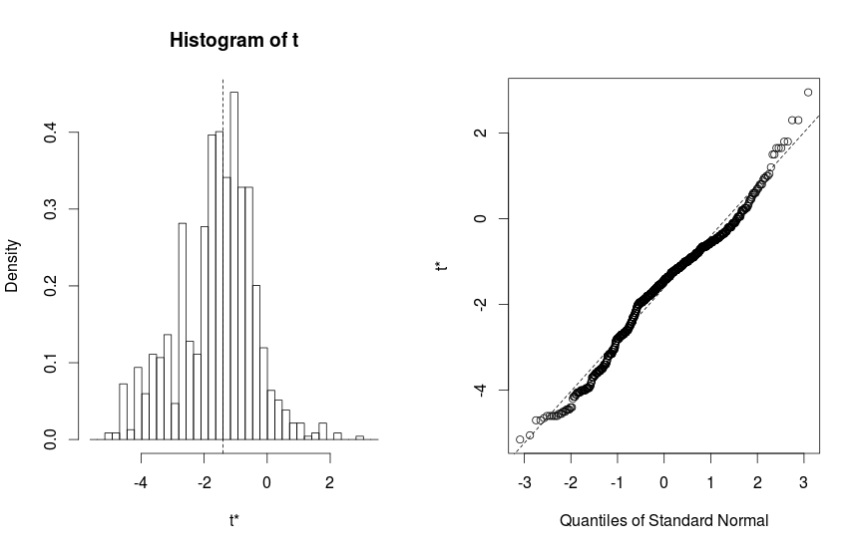
From the results, the difference in median number of hours of sleep (median after drug 1 – median after drug 2) is -1.4 hours (95% CI -4.29 to 0.75 hours). Since the 95% CI arms cross 0, we can conclude that when subjects took drug 1, they did not have fewer median number of hours of sleep compared to when they took drug 2.
The figure plots a histogram of the bootstrapped differences of the medians from all 1000 generated samples (left), and a quantile-quantile plot (right) comparing the the bootstrapped differences of the medians on the y axis to quantiles from a Normal distribution on the x-axis. The bell shape of the histogram, and the appearance of the quantile-quantile plot that shows bootstrapped differences of the medians lie along the diagonal line, indicate that the bootstrapped values follow a Normal distribution.
Summary
We calculated a difference of the medians between conditions in our sample, and used a resampling technique called bootstrapping to calculate 95% CI about our estimate of the difference of the medians. The real value of bootstrapping is that it can be used to calculate confidence intervals about non-parametric statistics that don’t rely on probability distributions.
